“Crowded, Compartmentalized, Sticky, Spatially Inhomogeneous”
In college, I remember being blown away by a huge, physical map of metabolic pathways our Biochemistry professor once brought into class. It looked like this:
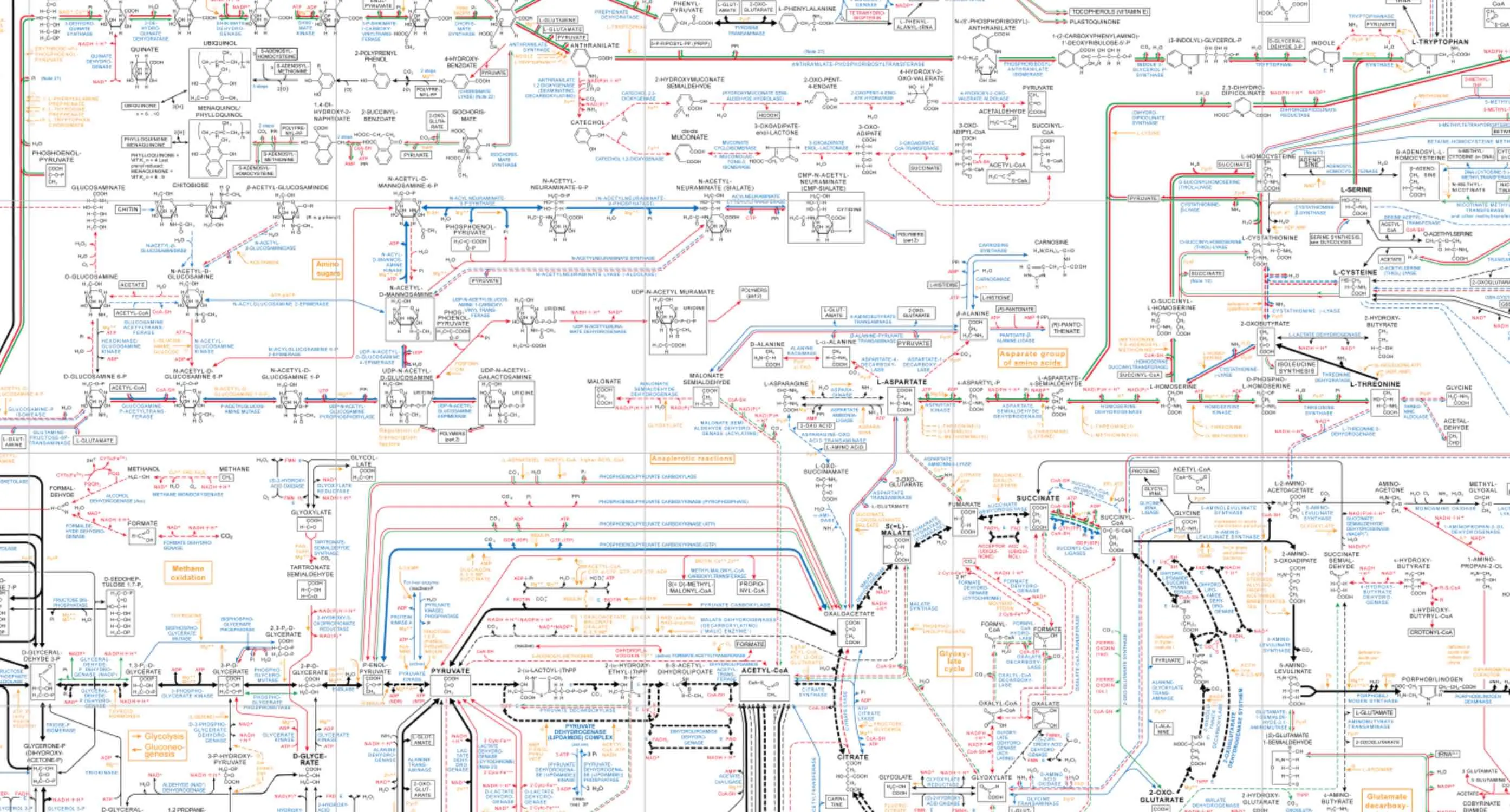
Here it is online. Kinda like a Google Maps of cellular reactions. It was impressed upon us that the interior of a cell (especially a eukaryotic one) is a really, really busy and tight and ‘goopy’ place: “crowded, compartmentalized, sticky, spatially inhomogeneous”. As that paper notes, this messy, “macromolecular crowding” helps your proteins fold properly (among several other factors.) This was a bit hard for me to appreciate since, up to then, I was only accustomed to images of cells from a light microscope or vastly simplified illustrations school texts.
I was somehow reminded of all of that after seeing some astounding paintings by Professor David S. Goodsell (Wiki, Twitter, Website). He calls the series “Molecular Landscapes.” Here are a few related to the pandemic we’ve been through.
SARS-CoV-2 and Neutralizing Antibodies, 2020
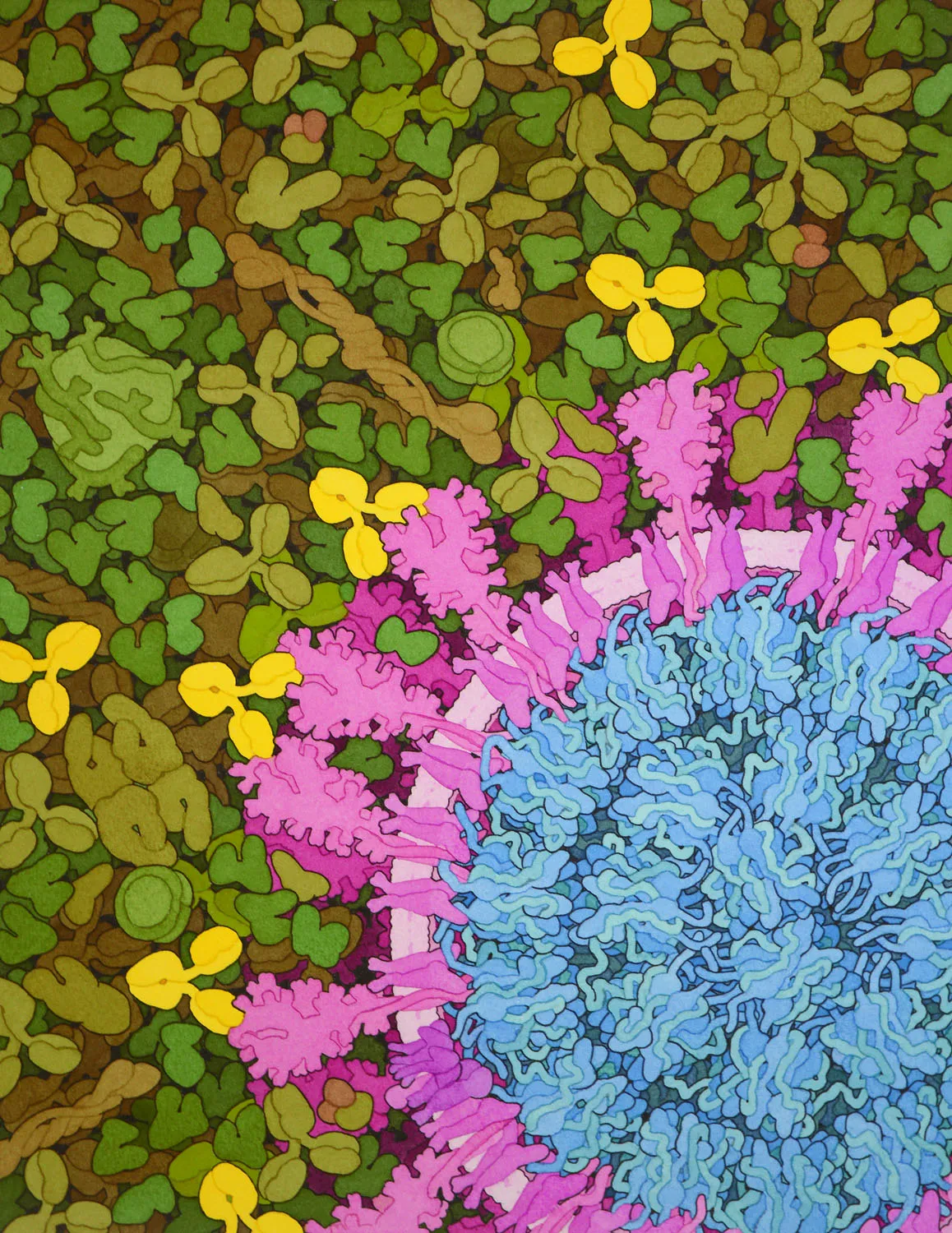

Acknowledgement: David S. Goodsell, RCSB Protein Data Bank and Springer Nature; doi: 10.2210/rcsb_pdb/goodsell-gallery-025. The painting was commissioned for the cover of a special COVID-19 issue of Nature, presented 20 August 2020, and is currently in the collection of the Cultural Programs of the National Academy of Sciences.
(Unknown Title)
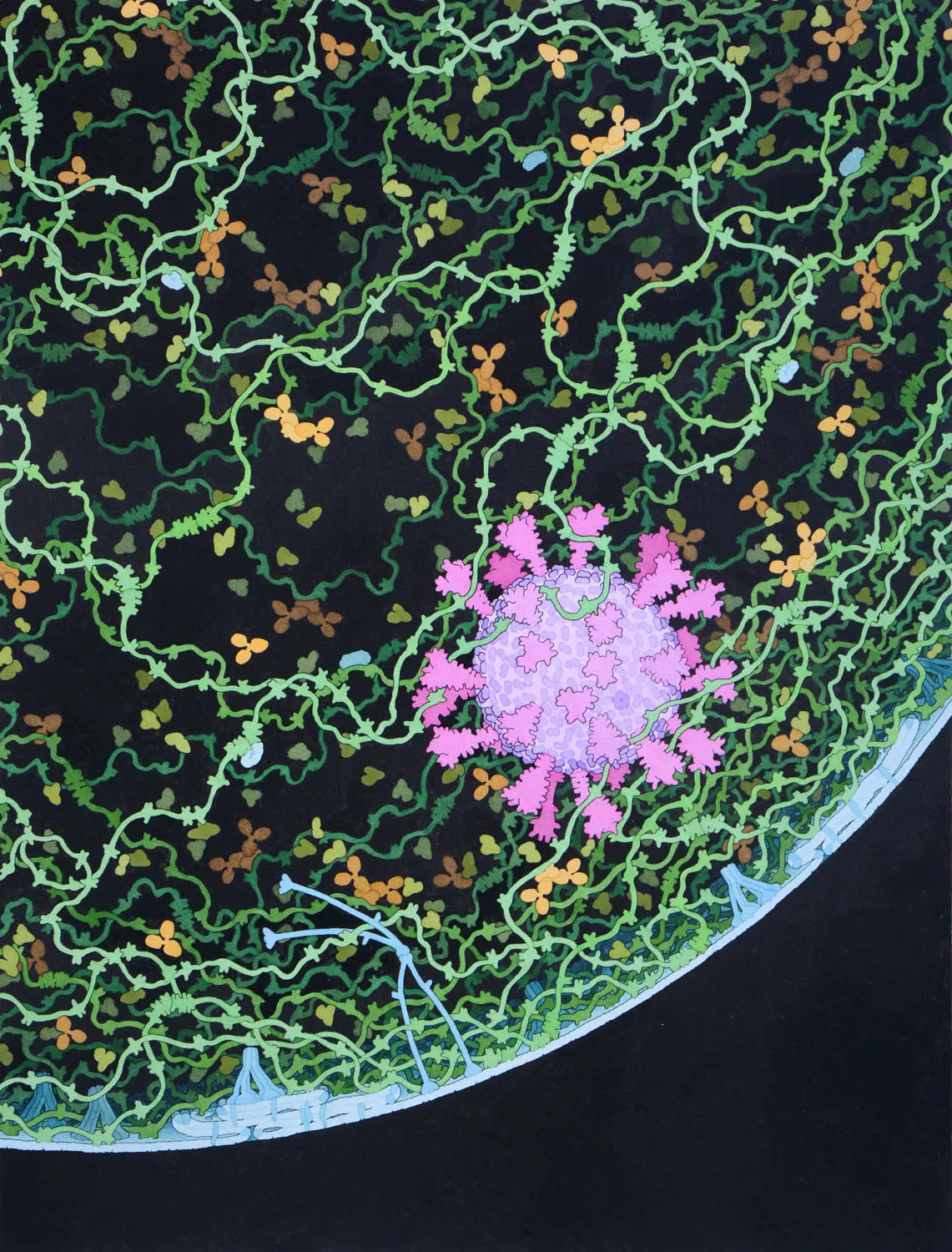
Acknowledgement: Illustration by David S. Goodsell
Coronavirus, 2020
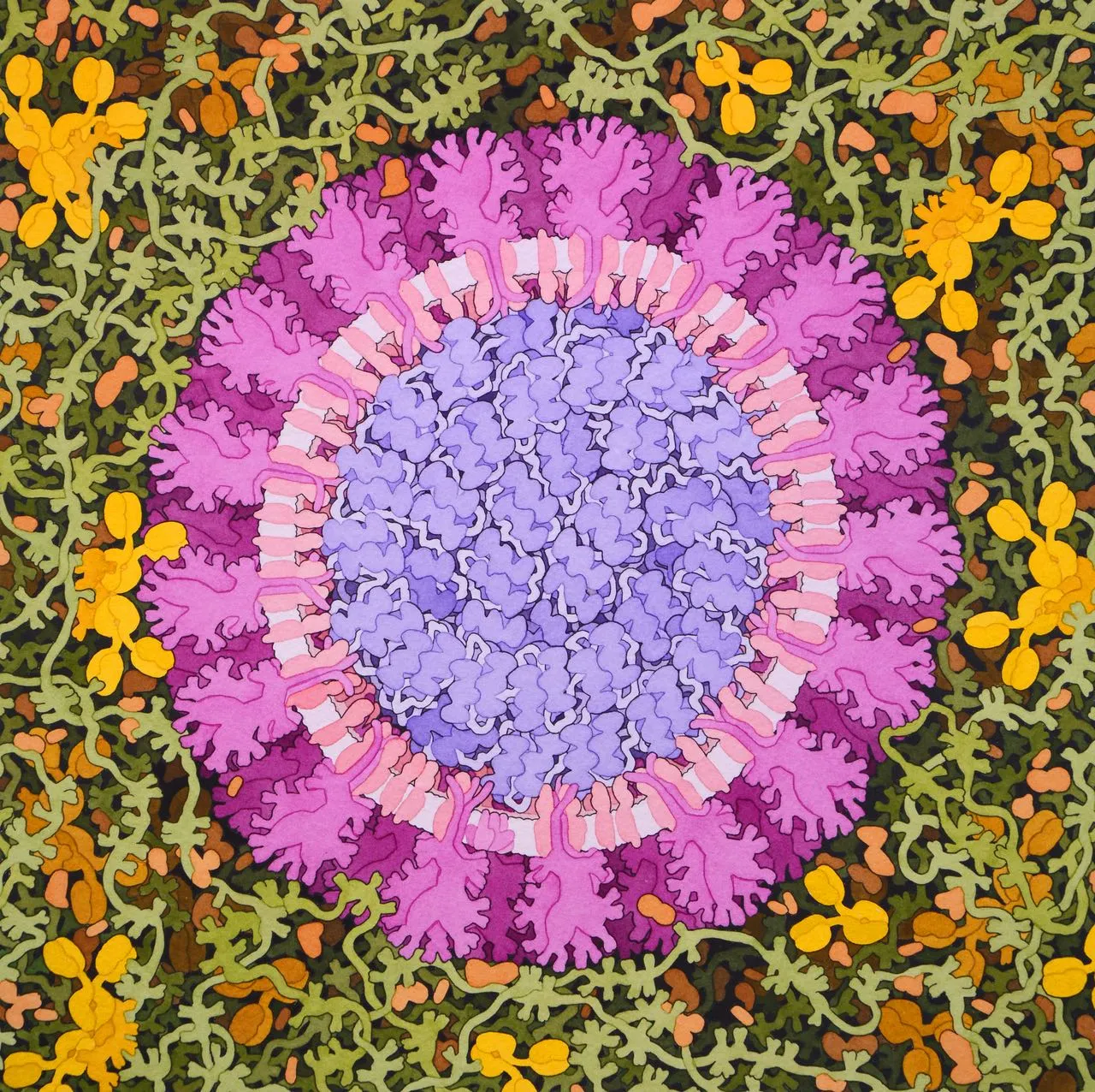

Acknowledgement: Illustration by David S. Goodsell, RCSB Protein Data Bank; doi: 10.2210/rcsb_pdb/goodsell-gallery-019. This painting depicts a coronavirus just entering the lungs, surrounded by mucus secreted by respiratory cells, secreted antibodies, and several small immune systems proteins. The virus is enclosed by a membrane that includes the S (spike) protein, which will mediate attachment and entry into cells, M (membrane) protein, which is involved in organization of the nucleoprotein inside, and E (envelope) protein, which is a membrane channel involved in budding of the virus and may be incorporated into the virion during that process. The nucleoprotein inside includes many copies of the N (nucleocapsid) protein bound to the genomic RNA.
SARS-CoV-2 Fusion, 2020
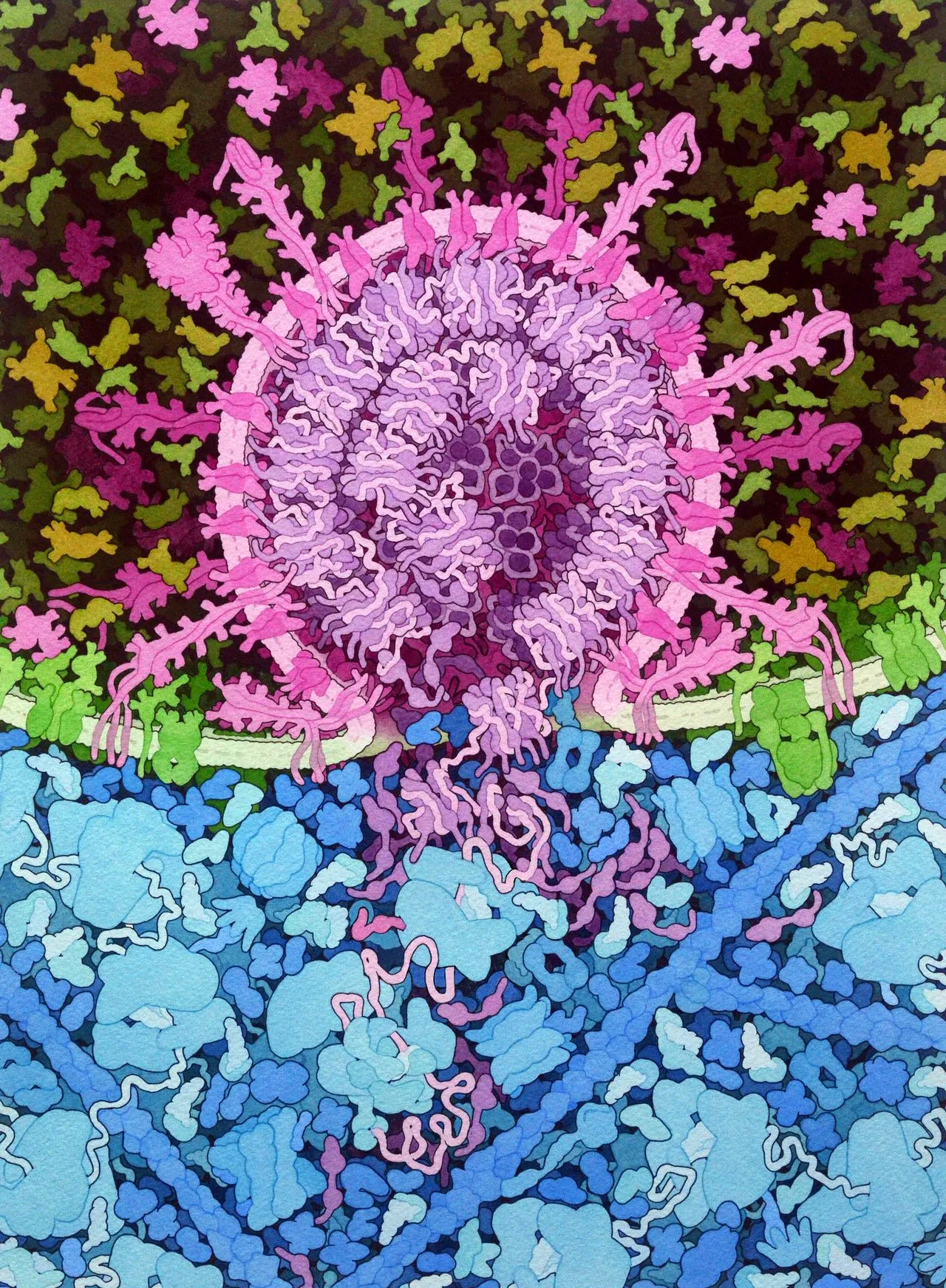
Acknowledgement: Illustration by David S. Goodsell, RCSB Protein Data Bank; doi: 10.2210/rcsb_pdb/goodsell-gallery-026. This painting depicts the fusion of SARS-CoV-2 (magenta) with an endosomal membrane (green), releasing the viral RNA genome into the cell cytoplasm (blue), where it is beginning to be translated by cellular ribosomes to create viral polyproteins. The painting includes speculative elements that are designed to highlight the process, most notably, multiple states of the viral spike protein are shown.
SARS-CoV-2 mRNA Vaccine, 2020
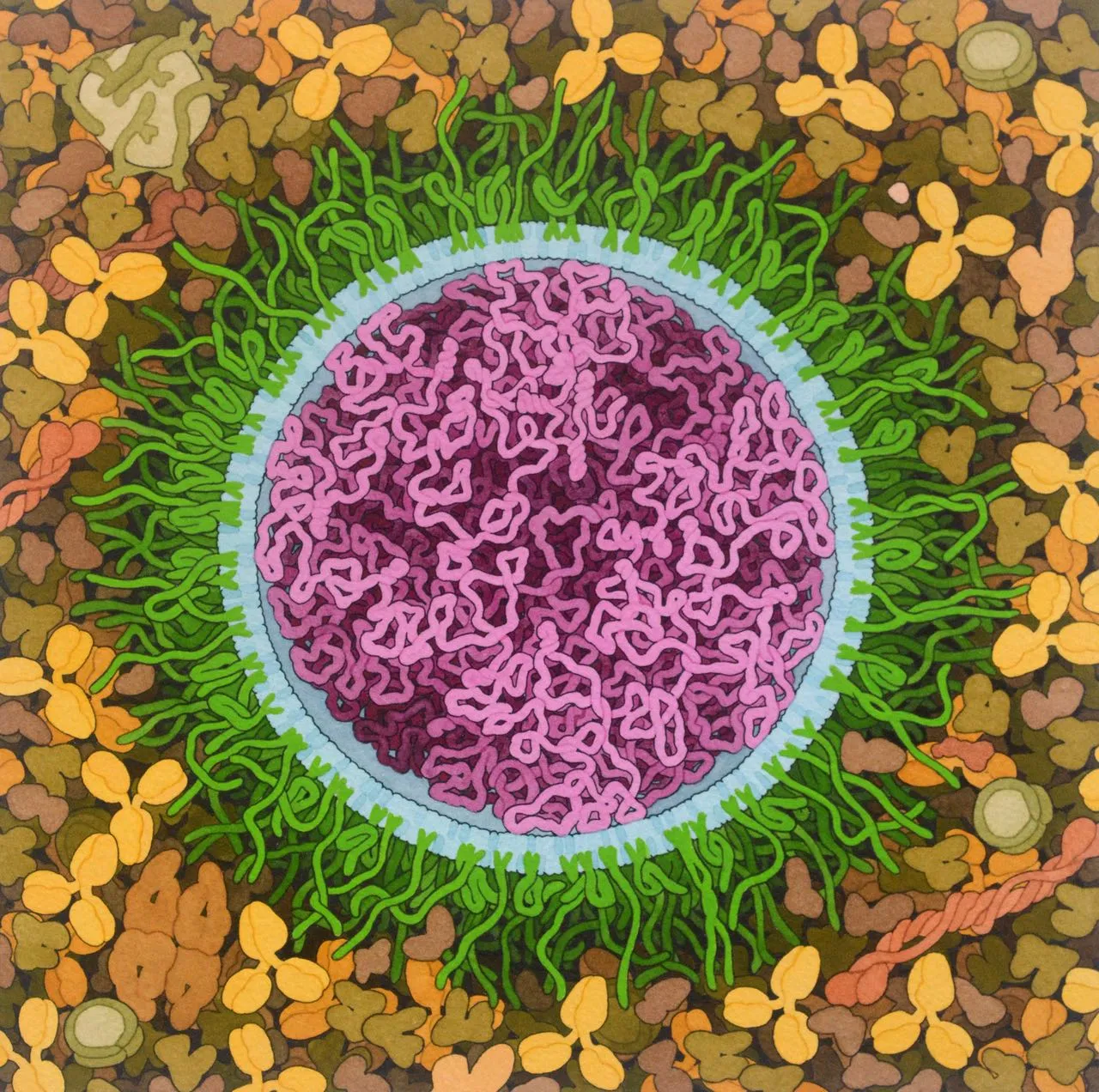

Acknowledgement: Illustration by David S. Goodsell, RCSB Protein Data Bank; doi: 10.2210/rcsb_pdb/goodsell-gallery-027. Messenger RNA (mRNA) vaccines developed for the COVID-19 pandemic are composed of long strands of RNA (magenta) that encode the SARS-CoV-2 spike surface glycoprotein enclosed in lipids (blue) that deliver the RNA into cells. Several different types of lipids are used, including familar lipids, cholesterol, ionizable lipids that interact with RNA, and lipids connected to polyethylene glycol chains (green) that help shield the vaccine from the immune system, lengthening its lifetime following administration. In this idealized illustration, all of the lipids are arranged in a simple circular bilayer that surrounds the mRNA and the PEG strands have both extended and folded conformations. In reality, the structure may be less regular, as suggested in the NanoLetters paper […]